June 2, 2006
Northwestern Researchers Work on Regeneration of Limbs
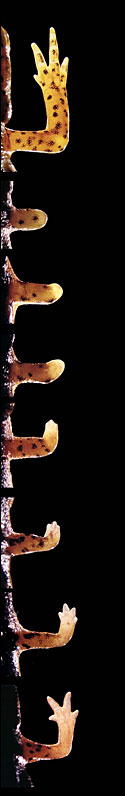 |
| Over six to eight weeks the newt can regenerate a limb after injury, as shown here in a sequence where the lighter color represents the newly formed forelimb. |
CHICAGO—Partial or complete loss of digits or limbs and deforming disabilities resulting from serious illness profoundly affect the quality of life of the wounded and remain a challenge for the medical community faced with treating them.
Recognizing the need for novel approaches that can restore, even partially, the structure and function of lost or damaged tissues, the Defense Advanced Research Projects Agency (DARPA) has awarded a $3.7 million grant to a consortium of six universities and research centers, including Northwestern University, to unlock the regenerative potential in humans.
Hans-Georg Simon, PhD, assistant professor of pediatrics at the Feinberg School of Medicine and a leading researcher at Children’s Memorial Research Center, heads the Northwestern University arm of the study. The DARPA grant will be administered through the University of Pittsburgh McGowan Institute for Regenerative Medicine.
The diverse group of researchers hopes that by working together they will gain a more complete understanding of the cellular and molecular basis that allows certain creatures, such as salamanders, to completely regenerate lost limbs and be able to harness this capacity in mammals.
If successful and measurable progress is made along the way, DARPA could provide the researchers up to $15 million in funding over four years.
“I don’t think it’s complete fantasy,” said Dr. Simon, who studies salamander regeneration at Children’s Memorial Research Center.
“The human body has quite remarkable capabilities for repair and regeneration. The problem is that we tend to lose the capacity as we age,” Dr. Simon said.
Most adult mammals, including humans, exhibit only limited regenerative abilities, and following an injury wounds usually heal over by forming a scar. However, evidence exists for regeneration-like processes during mammalian embryonic development, and lifelong self-renewal capability is evident for selected cell populations such as blood cells, cells that line the walls of the intestine, and liver cells.
Reports indicate that the tip of the finger can sometimes be regenerated if the cut is above the last joint, Dr. Simon said.
The team of individual research groups tries to use a solution that nature itself has developed for repairing damaged limbs or organs in a wide variety of animals. Many species including certain salamanders and zebrafish can regenerate a wide variety of body parts. The salamander can regenerate its limbs, tail, spinal cord, upper and lower jaws, and intestine, part of the heart, and the lens and retina of the eye. The zebrafish can regrow its fins, scales, and spinal cord and part of the heart.
“Regeneration is the most complete repair mechanism there is,” noted Dr. Simon.
Dr. Simon’s laboratory group successfully identified genes that control the specific shape of developing forelimbs (arms) and hindlimbs (legs) in virtually all vertebrates, from fish to birds to mice to humans. They discovered that these genes also play a critical role in heart development and that mutations in humans result in birth defects displaying malformations in both the limbs and the heart.
In other research Dr. Simon and Northwestern scientist Stuart R. Stock, PhD, research professor of molecular pharmacology and biological chemistry, used digital x-ray microtomography to discover the ways in which salamanders form bone and cartilage during limb regeneration.
“Regenerating salamander cells are similar to activated stem cells. However what makes them special is that they contain a complete blueprint of the biological structure they have to rebuild. The clinical benefits of developing similar tissue regeneration in humans are obvious, but even in regenerating species such as the salamanders, the basic biology of regeneration is not fully understood,” Dr. Simon said.
The regenerative processes are dependent upon the formation of a blastema, a mass of immature and unspecialized cells, at the site of injury. The regeneration blastema consists of precursor cells originating by local de-differentiation of adult stump tissue. In de-differentiation, specialized cells or tissues revert to a simpler, more embryonic, unspecialized form.
The blastema performs the difficult task of integrating new and existing tissues. The blastema is a self-organizing system, suggesting that autonomous gene pathways control both tissue and position-specific developmental programs.
To gain an understanding of the self-regulating mechanisms and pathways operating within the blastema, the Simon laboratory is developing a regeneration gene array to explore the unique gene activities in regenerating salamander forelimb, hindlimb, and tail tissues.
By comparing the expression patterns of these genes between the different appendages, they try to identify a set of genes that are consistently upregulated in the regenerating limbs and tail. Upregulation is the process by which a cell increases the number of active genes to achieve a certain function.
These gene activities will be compared with data from team member Ellen Heber-Katz, PhD, a researcher at the Wistar Institute in Philadelphia, who is performing similar studies with a special “super-healer” or MRL mouse strain.
“This will give us insight into the types of molecular events that are controlling the regenerative response across species and will allow us to identify a signature of gene activities that are important for tissue regeneration,” Dr. Simon said. “This is still a complicated story, but we have the first inroads now. I think we’re getting into a very exciting time.”
In addition to Drs. Simon and Heber-Katz, members of the research team are Stephen Badylak, PhD, University of Pittsburgh McGowan Institute; Lorraine Gudas, PhD, Weill Medical College of Cornell University in New York; Shannon Odelberg, PhD, University of Utah, Salt Lake City; and Susan Braunhut, PhD, and Kenneth Marx, PhD, University of Massachusetts at Lowell.